Imagine you’re standing on the edge of a lake, lined with beautiful flowers and high grass. There is a shallow part right in front of you populated with reed, birds and frogs basking in the sunlight. You have a rock in your hand the size of a golf ball and without a good reason you throw it as far as you can. The rock hits the water with a splash, creating a perfect circular rippling effect. While the disturbance grows larger, the ripples are dampened, and finally the disturbance disappears. Out of nowhere a helicopter appears with an enormous boulder tethered underneath. The boulder gets dropped into the same lake from 300 feet, and creates a circular wave that carries across the entire lake and seriously disrupts the surrounding wildlife. If you hadn’t seen the rock drop in the lake, you could have guessed from the breadth of the disturbance that the initial shock to the system must have been significant. This simple thought-experiment formed the basis behind our contemplated approach on how an antibiotic may disturb and affect a bacterium, which resulted in a new biological concept that defines the level of stress a bacterium is experiencing independent of strain, species or type of stress.
So far, several approaches have been developed that use the transcriptional response of a small number of genes (a gene-panel) to determine whether a bacterium is sensitive or resistant to a specific antibiotic. It turns out that this can work pretty well. For instance, the expression profile of a set of 5 genes can indicate whether E. coli is sensitive or resistant to ciprofloxacin (a fluoroquinolone antibiotic). This information can be quite useful when making treatment decisions for an E. coli strain that is causing an infection in a patient. However, we show that there are several issues with such an approach: 1. Not all bacterial strains and species respond to an antibiotic in the same way and with the same set of genes; 2. Genes that are indicative of a response in one species may be poorly conserved across strains and species; 3. There is not always a clear rationale of how genes are selected for a gene-panel. These and several other factors prevent the generalizability and rapid implementation of such an approach. This means that every time you encounter a new strain or species you would need to generate a large dataset for each drug to determine which genes should be monitored to indicate drug-sensitivity.
In an attempt to design an approach that works for most any strain, species and stress-type (drug) we built on our ‘boulder in the lake’ analogy and wondered whether it is possible to measure the amount of stress that develops across an entire system while the system is being disturbed. We reasoned that like the golf ball example, an antibiotic at a concentration below the minimum inhibitory concentration (MIC) would create localized stress that eventually gets dampened. In contrast, an antibiotic exceeding the MIC would, like the boulder dropped from the helicopter, carry throughout, and disturb the entire system. In turn this made us think of how chaos can develop in a system over time, which can be captured by quantifying the amount of entropy. To explore this idea, we generated a large set of RNA-Seq profiles for over 10 different bacterial strains and species, we evolved different bacteria to become resistant to different antibiotics or grow in the absence of formerly essential nutrients.
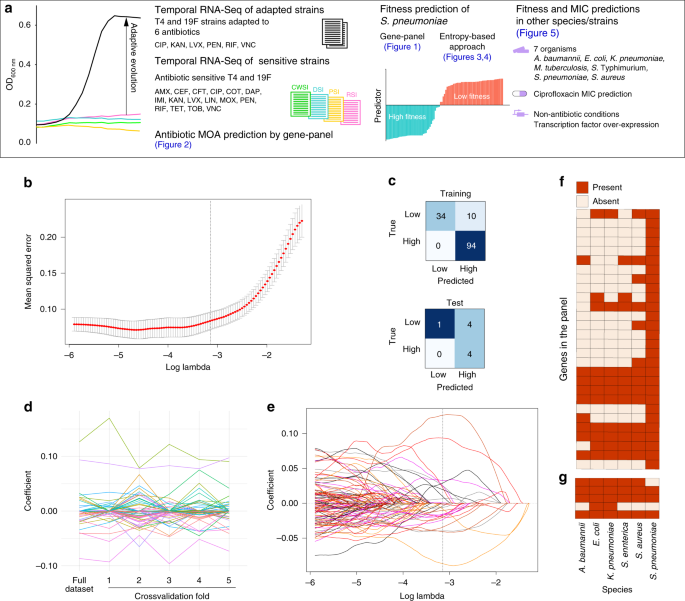
We combined all of these experiments and data, as well as several published data sets, and designed an approach that measures transcriptional entropy across the entire genome, and implements it as a defining building block in a set of models that make highly accurate predictions on antibiotic sensitivity. For instance, we show that every class of antibiotics has its own type of response, and some antibiotics trigger rapid responses, while other responses are slower. With increasing antibiotic concentration, a response carries further through the genome with increasing chaos/entropy as a result. And finally, when entropy increases beyond a specific threshold, which becomes clear within 30-90 minutes after the start of the stress-exposure, the bacterium is ‘doomed’ and will not be able to withstand the stress. We show that there are a variety of ways to calculate and implement entropy, from a fairly complex approach by using temporal measurements and genetic relationships between all the genes in the genome, to a simple approach where only a single measurement is needed. Moreover, we show that entropy is not just a binary indicator of stress, instead the level of entropy is directly related to the MIC of the bacterium. This means that by exposing a bacterium to a single antibiotic concentration and taking a single time measurement, the level of chaos/entropy accurately predicts the antibiotic’s MIC.
The manuscript lists other findings and new approaches we developed along the way, including an antibiotic’s mechanism of action predictor and an approach to extract gene-dependencies. Most importantly, we show that transcriptional entropy makes generalizable stress predictions possible that are independent of strain, species or type of stress and can even be implemented to make quantitative predictions. We envision this work opens up exciting new avenues, including in vivo stress-response profiling to predict how both the host and the pathogen is developing and whether for instance treatment is necessary, which could propel entropy-based predictions into a clinical setting.
Zhu Z.*, Surujon D.*, Ortiz-Marquez J.C.., Huo W., Isberg R. R., Bento J., and van Opijnen T. 2020 Entropy of a bacterial stress response is a generalizable predictor for fitness and antibiotic sensitivity. Nature Communications 11, 4365. doi: https://doi.org/10.1038/s41467-020-18134-z




Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in