In September 2015, the United Nations adopted 17 major sustainable development goals to address global challenges and build a more sustainable future. Among them are “Zero Hunger” and “Responsible Consumption and Production”, which are becoming increasingly difficult to achieve with continued population growth. Finding solutions to this, while reducing greenhouse gas emissions and protecting valuable ecosystems, remains a defining challenge of our era. An area where such solutions are urgently needed is in food loss, where the microbial spoilage of food is estimated to account for one-fourth of global food waste. Despite increasing efforts by the food industry in the last decades, food products regularly become contaminated with pathogenic or spoilage bacteria during primary processing. While efforts to date have been focused on different disinfection strategies and the detection of specific pathogens, little is known about the microbial diversity residing in food processing plants. Even less is known about how these microbes migrate throughout the facilities and which migration routes lead to food contamination.
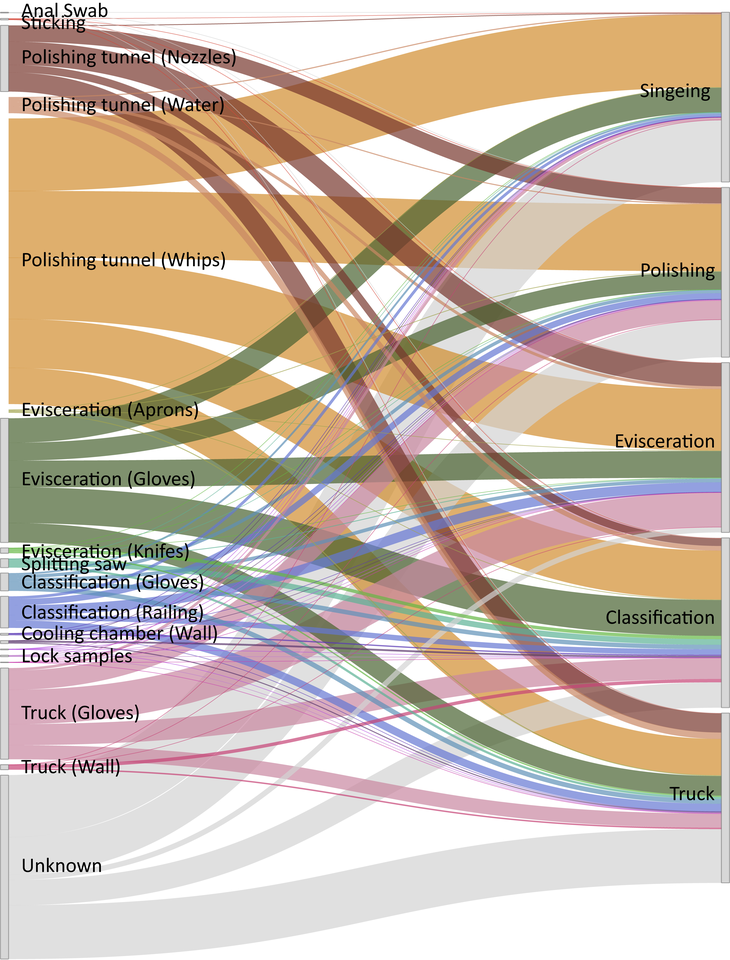
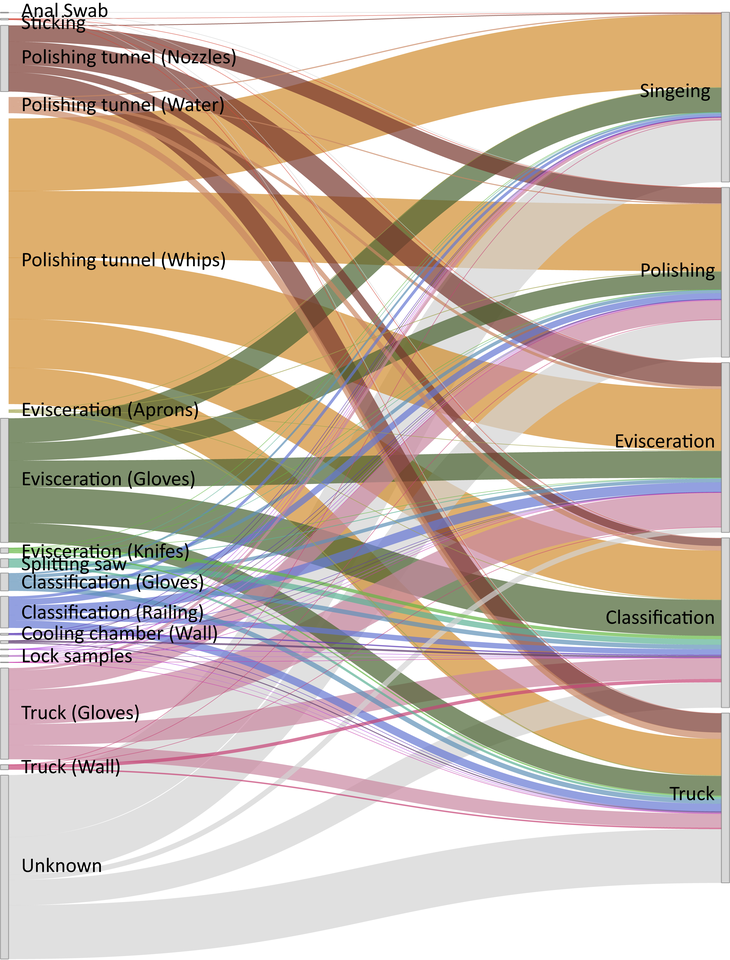
When I started my PhD at the University of Veterinary Medicine in Vienna in 2017, I joined a project that aimed to study the transmission patterns of microbes along the meat production chain. It soon became clear, due to limitations in throughput and culturability, that we would not be able to track the majority of microbes with traditional microbiological techniques. We therefore turned to next generation sequencing (NGS) techniques, which have been broadly applied in characterizing microbial communities from diverse habitats. At the time, Pacific Biosciences had just released the PacBio Sequel system, a platform that enables sequencing of hundreds of thousands of highly accurate long reads in a single run. Using this platform, we were able to detect exact amplicon sequence variants that represented the full-length 16S rRNA gene in high throughput. We then applied the software SourceTracker, which predicts the source of microbial communities in designated sink samples, to trace transmission routes of the sequence variants and propose sources of specific microbes. With these data we created a facility-specific transmission map of bacterial flow which predicted previously unknown sources of bacterial contamination during meat processing (Figure 1). This provided the facility with essential information for the targeted combat of potential pathogens or meat-spoilage organisms.

Continued advances in long-read sequencing technologies will further increase throughput and taxonomic resolution, enhancing its potential in regular food monitoring applications. However, the current application of such an approach requires highly skilled staff and lacks standardization. Simplifying and standardizing protocols, from sample to bioinformatic analysis, will eventually allow for the regular use of NGS in food production facilities, the direct comparison of datasets and ultimately a better understanding of microbial contamination along the food supply chain.

Please sign in or register for FREE
If you are a registered user on Research Communities by Springer Nature, please sign in